Certain Bacterial Strains Are Present in Different Amounts Between Acne and Non-acne Patients, and These Strains Display Varying Effects on the Immune System, Which Could Affect Acne Development Differently

The Essential Info
Acne is in part a bacterial disease. When it comes to acne, a specific type called C. acnes is the culprit. Scientists found that C. acnes may:
- Contribute to the formation of clogged pores
- Cause irritation inside clogged pores once they are formed
- Cause pimples to become more inflamed, red, and sore
This article is one of the deep science type articles here on Acne.org. Read on if you’re Team Nerd: There are multiple C. acnes strains which are present in different amounts in the skin of acne-prone people. For example, C. acnes type IA1 is present in higher amounts in acne-prone skin than in healthy skin. Further, preliminary research suggests that this strain demonstrates a slightly altered effect on the immune system when compared with other types, and consequently likely contributes to acne development differently.
Research in this area is still in its beginning stages, and scientists will need to perform further research to better characterize the differences among the types of C. acnes as well as other bacterial species, such as Staphylococcus and Klebsiella, to determine how each affects acne development.

The Science
- Role of C. acnes in Acne Development
- Bacterial Strains of C. acnes and Acne Development
- Other Bacterial Species in Acne Development
Normally, I’ll try to make articles as understandable as possible. I’ll do that with this one as well, but this is deep science type stuff, so let’s throw caution to the wind and just delve right into all the gory details. Put your thinking cap on and let’s do this!
Many factors lead to the formation of acne lesions, including:
- Overproduction of skin oil, called sebum
- Skin cell overgrowth
- Hormones
- Bacteria
For the purpose of this article, we’ll focus in on bacteria. It turns out that acne-prone and healthy individuals carry different bacteria and bacterial strains in their skin, only some of which may contribute to the formation of acne. The hope is that if we can discern the exact type of bacteria that is leading to acne symptoms, we can then target it better with treatments.
First, let’s look at the role of acne bacteria, called C. acnes, in acne development.
Role of C. acnes in Acne Development
Cutibacterium acnes (C. acnes) is a single-cell bacterium that is present in both healthy skin and acne-prone skin. It survives using sebum as a source of nutrients, and therefore thrives in clogged pores, where sebum is abundant.
Although scientists previously thought C. acnes was the main cause of acne, recent research found that it is not present in all acne lesions, and therefore is likely to play only a secondary role in acne formation.1 In other words, C. acnes doesn’t cause acne, but it can make acne worse.
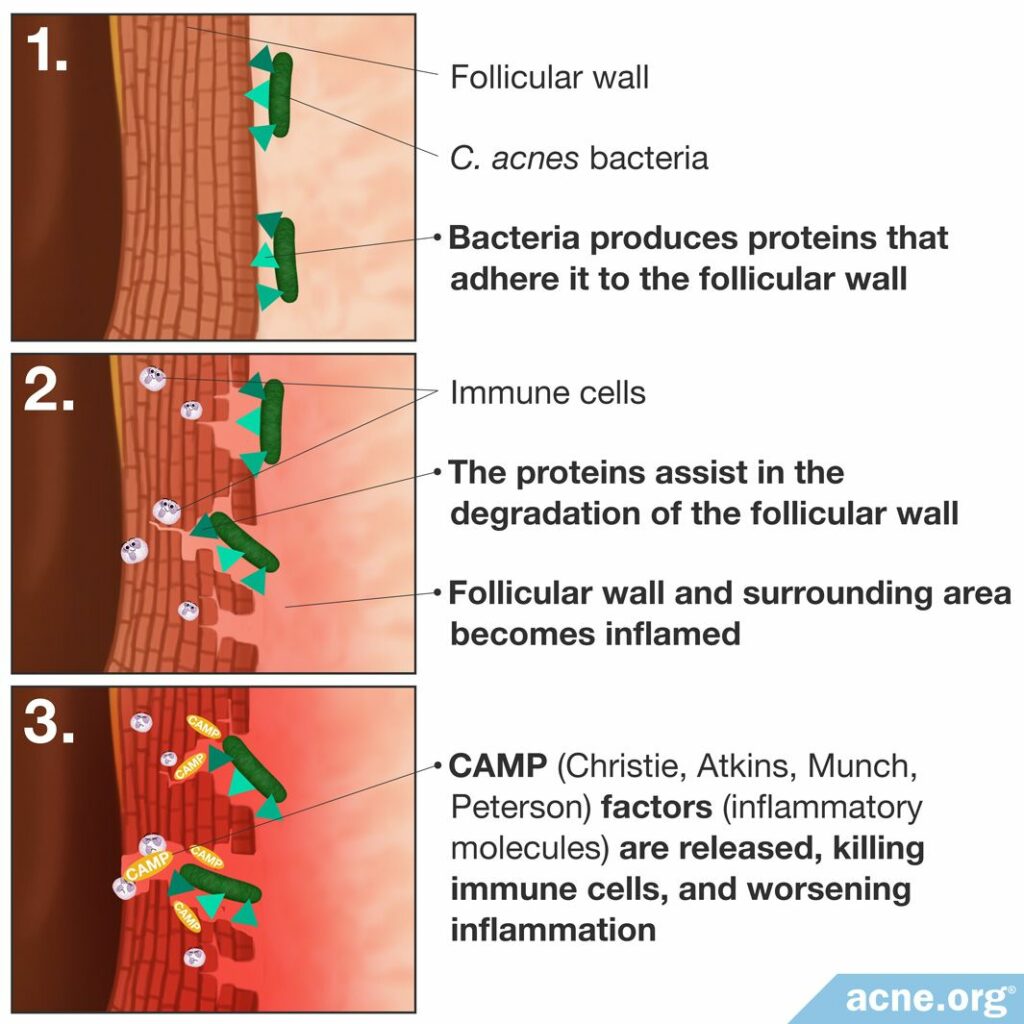
Researchers now believe that C. acnes acts to contribute to the worsening of previously formed acne lesions in three main ways.
- Producing proteins that adhere to the pore wall. Once the bacterium is stuck to the pore wall, it can better survive and reproduce.2,3
- Creating and releasing certain proteins that assist in the degradation of the pore wall, contributing to its rupture and ensuing inflammation.2,3
- Stimulating the immune response by triggering the release of inflammatory molecules, which results in inflammation. One main class of inflammatory molecules that C. acnes releases is called CAMP (Christie, Atkins, Munch, Peterson) factors. CAMP factors are able to puncture immune cells, resulting in immune cell death, toxicity, and release of additional inflammatory molecules that worsen inflammation.2-5
Bacterial Strains of C. acnes and Acne Development
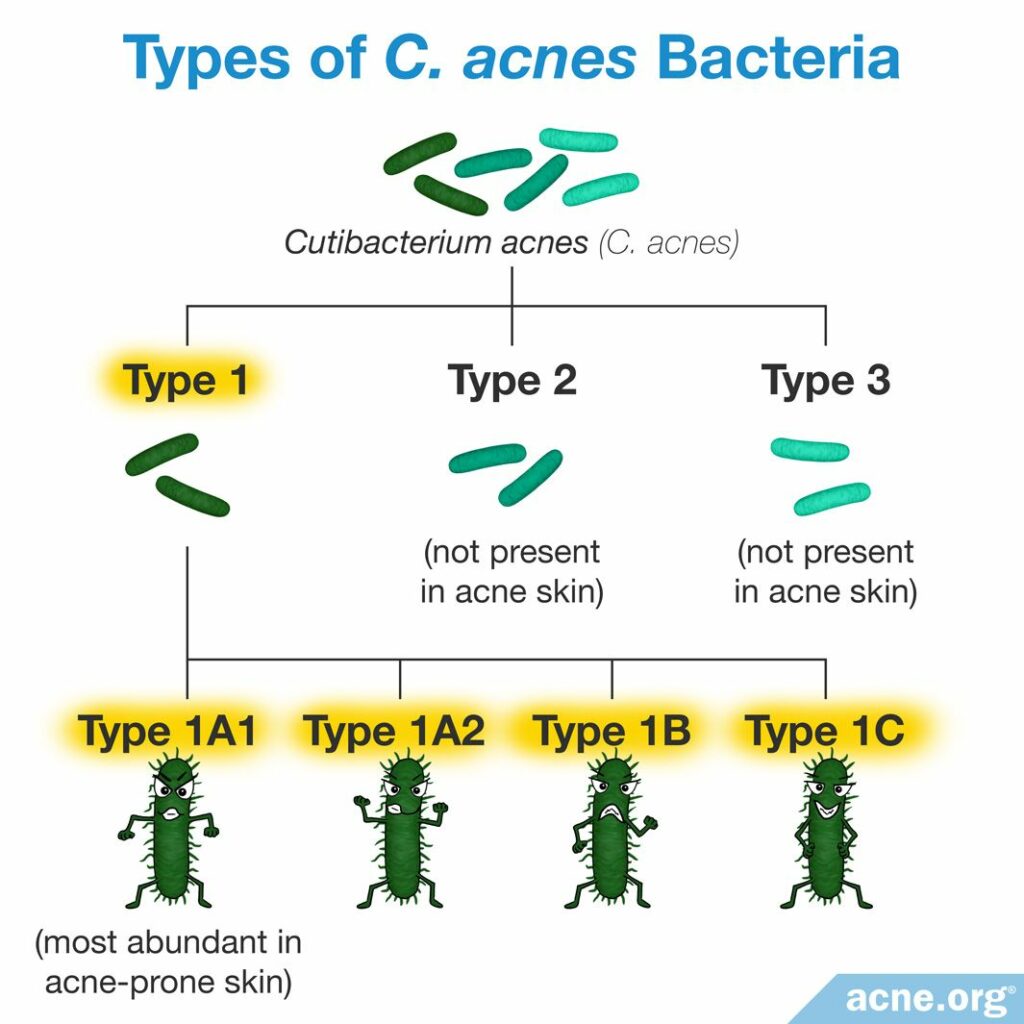
Scientists classify bacteria of the same species into small subgroups called strains. To group bacteria into strains, scientists sequence the bacterial genome and group bacteria into strains if they have at least 95% identical DNA. There are about 100 strains of C. acnes that fall into the following 3 main groups:
- Type I – present in skin with acne
- Type II – not present in skin with acne
- Type III – not present in skin with acne
Type I contains the additional 4 strains:6
- Type IA1 – most abundant in acne-prone skin
- Type IA2
- Type IB
- Type IC
Scientists have performed 4 independent studies to identify if the different strains present in the skin influence acne development differently. All 4 studies found that type IA1 is more abundant in acne-prone skin than in healthy skin.7-10
Further, one of the studies found that type II bacteria were not present in skin samples taken from skin with acne, and another study showed that the same was true of type III bacteria. But in both studies, type II and type III bacteria made up around 20% of all bacteria in the healthy skin samples.8,10
With the knowledge that type IA1 is more abundant in acne-prone skin than in healthy skin, scientists have performed four studies to classify if this particular strain of C. acnes contributes to acne development in a unique way.
Results of these studies show that type IA1 may be better able to survive inside clogged pores, which explains why we see them more often in acne-prone skin. Also, type IA1 may provoke the immune system in different ways than the other strains, but researchers need to perform more research to understand exactly how this might affect acne development.
Expand to read details of studies

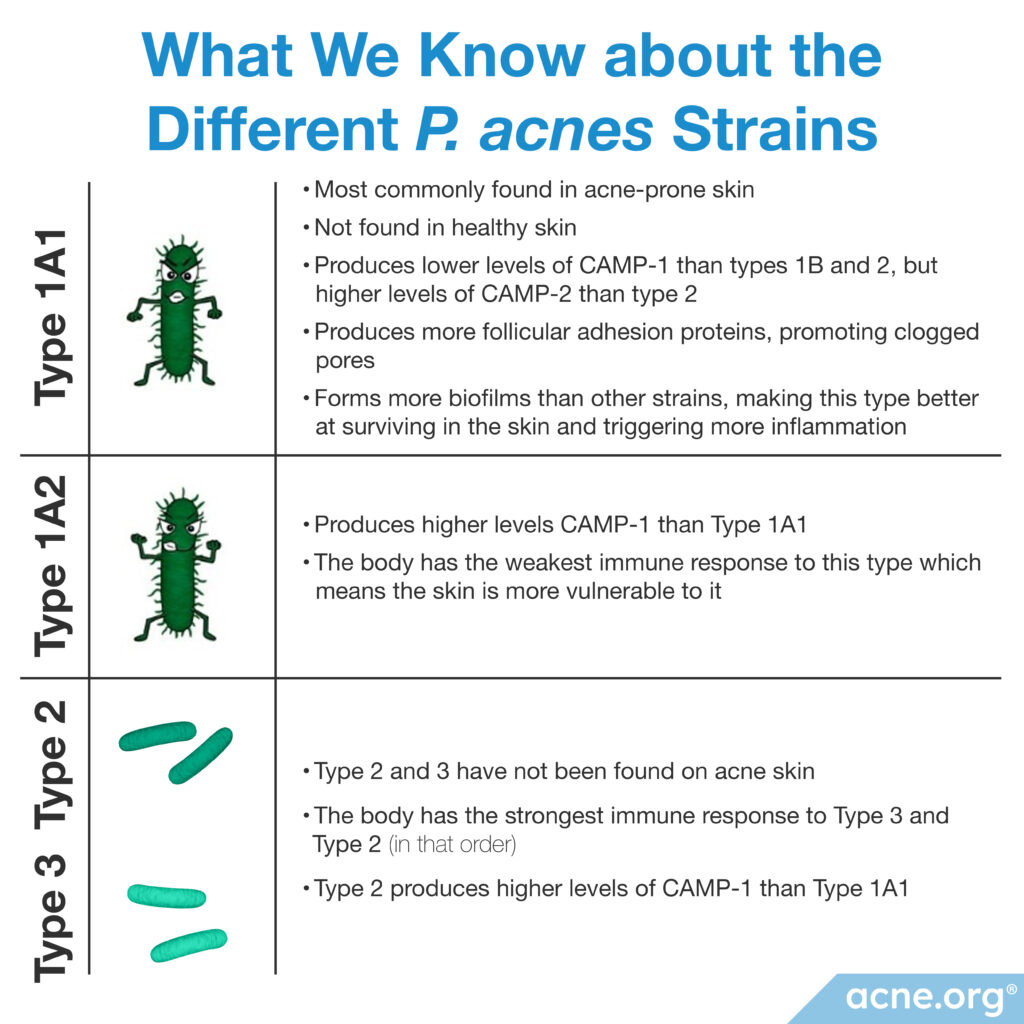
A 2005 study published in the journal Microbiology found that C. acnes type IA1 produced much lower levels of CAMP-1 than types IB and II, but higher levels of CAMP-2 than type II. Although it remains unclear how this affects the inflammatory response, this study concluded that different C. acnes types could provoke unique immune responses.11

A 2010 study published in BMC Microbiology discovered that C. acnes type IA1 produced more follicular adhesion proteins, which likely helps the bacteria stick better to the follicular (pore) wall, thus promoting clogged pores.12

A 2013 study published in Experimental Dermatology examined the immune system response to C. acnes in laboratory-grown human cells. They used their data to order the C. acnes types by strongest immune response (type III) to weakest immune response (type IB). They found that the order of immune system response from strongest to weakest is type III, II, IC, IA1, and IB. Further, the scientists found that each C. acnes type stimulated the release of different inflammatory molecules called cytokines, which in theory could stimulate unique immune system responses.13

A 2018 study published in the International Journal of Medical Microbiology looked at the ability of different C. acnes strains to form biofilms.14 A biofilm is a large cluster of bacteria embedded into a sugar matrix produced by the bacteria themselves. Bacteria living in biofilms are more resistant to antibiotics and more likely to trigger inflammation than bacteria living on their own. Recent research also suggests that C. acnes biofilms produce substances that change the composition of skin oil. This seems to create a vicious cycle, because once the composition of skin oil changes, this makes the skin oil an even more attractive environment for C. acnes bacteria to live in.15 Research shows that people with acne tend to have significantly more C. acnes biofilms in their skin pores than people without acne.15,16
This study found that C. acnes type IA1 bacteria were 2-8 times better at forming biofilms compared to other strains.14 Since biofilms help acne bacteria to survive and to stimulate more inflammation, this may make type IA1 bacteria especially problematic for people with acne.
Other Bacterial Species in Acne Development
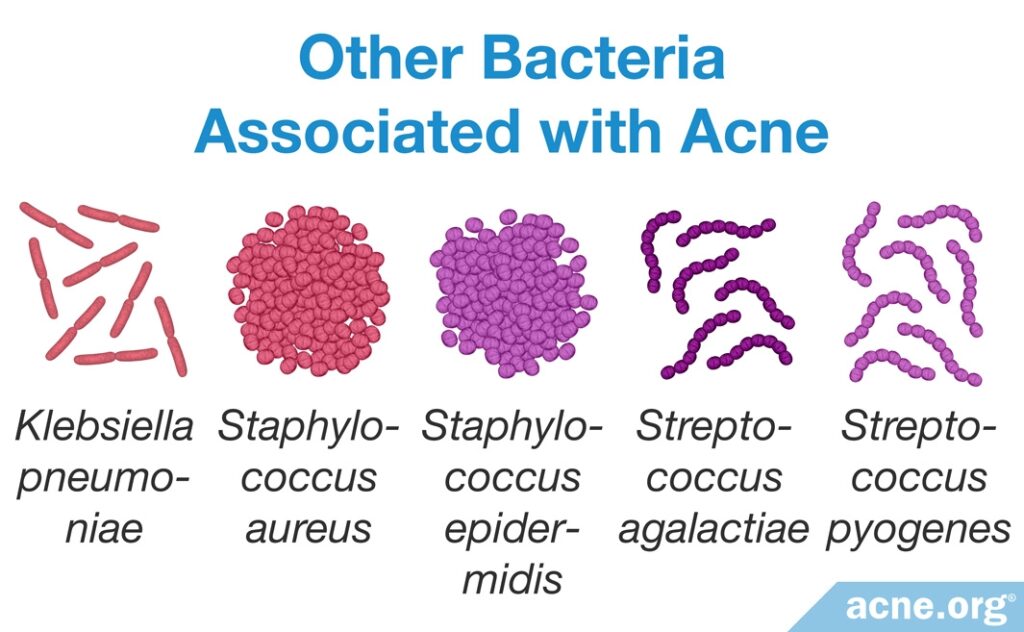
Although scientists have focused their acne development research on C. acnes, there is a variety of other bacterial species present in the skin that also may affect acne formation.
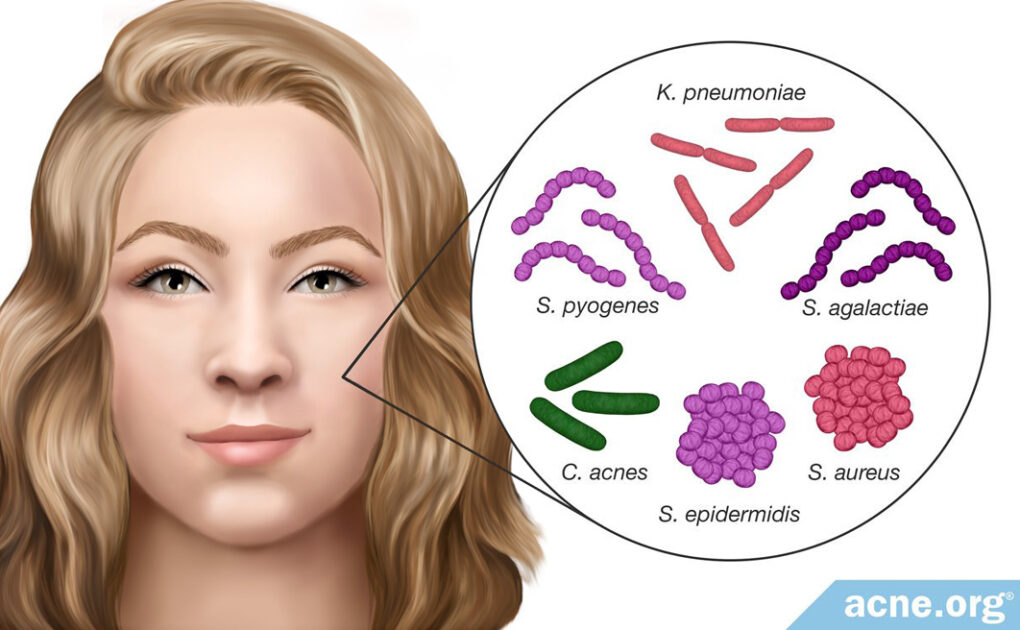
- Researchers found that the presence of Staphylococcus epidermidis (S. epidermidis) is increased by 70% in acne-prone skin. S. epidermidis might be overgrowing in skin pores and even crowding out C.acnes, resulting in an imbalance of skin bacteria.17,18 Further, research shows that S. epidermidis secretes factors that can worsen pore clogging.
- It also was noted that Staphylococcus aureus (S. aureus) is present at higher levels in many skin disorders, including acne. Scientists believe that S. aureus may produce proteins that aid in pore wall rupture.
- In addition, researchers found Staphylococcus pyogenes (S. pyogenes) in the oropharynx (region including the back of the throat and tongue) in acne-prone individuals but not healthy individuals. It is not yet clear how bacteria present in the oropharynx affects the development of acne.
- Klebsiella pneumoniae (K. pneumoniae) also was found in acne lesions, but not in healthy skin.
- Streptococcus agalactiae (S. agalactiae) produces factors that are similar to those of C. acnes and may be involved in acne development.1
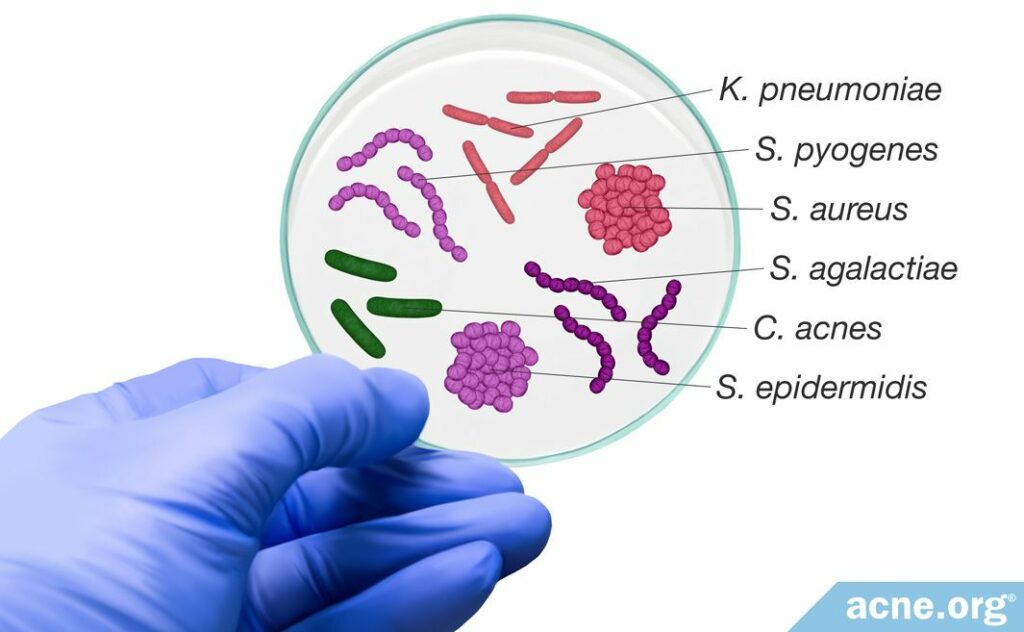
The research into the specifics of various bacteria strains is an exciting area of exploration. However, it is just getting underway. Scientists have performed scarce research examining the various roles these bacteria play in acne development, and therefore researchers must perform more studies to confirm these associations.
References
- Shaheen, B. & Gonzalez, M. A microbial aetiology of acne: what is the evidence? Br J Dermatol 165, 474 – 485 (2011). https://www.ncbi.nlm.nih.gov/pubmed/21495996
- Kumar, B. et al. New insights into acne pathogenesis: Exploring the role of acne-associated microbial populations. Dermatol Sinica 34, 67 – 73 (2016). https://www.sciencedirect.com/science/article/pii/S1027811715001457
- Brüggemann, H. et al. The complete genome sequence of Propionibacterium acnes, a commensal of human skin. Science 305, 671 – 673 (2004). https://www.ncbi.nlm.nih.gov/pubmed/15286373
- Nakatsuji, T., Tang, D. C., Zhang, L., Gallo, R. L. & Huang, C. M. Propionibacterium acnes CAMP factor and host acid sphingomyelinase contribute to bacterial virulence: Potential targets for inflammatory acne treatment. PLoS ONE 6, e14797 (2011). https://www.ncbi.nlm.nih.gov/pubmed/21533261
- Dréno, B., Dagnelie, M. A., Khammari, A. & Corvec, S. The skin microbiome: A new actor in inflammatory acne. Am J Clin Dermatol 21 Suppl 1, 18-24 (2020). https://pubmed.ncbi.nlm.nih.gov/32910436/
- Yu, Y. et al. Typing of Propionibacterium acnes: a review of methods and comparative analysis. Br J Dermatol 172, 1204 – 1209 (2015). https://www.ncbi.nlm.nih.gov/pubmed/25600912
- Lomholt, H. B. & Kilian, M. Population genetic analysis of Propionibacterium acnes identifies a subpopulation and epidemic clones associated with acne. PLoS ONE 5, e12277 (2010). https://www.ncbi.nlm.nih.gov/pubmed/20808860
- McDowell, A. et al. An expanded multilocus sequence typing scheme for Propionibacterium acnes: Investigation of ‘pathogenic’, ‘commensal’ and antibiotic tesistant strains. PLoS ONE 7, e41480 (2012). https://www.ncbi.nlm.nih.gov/pubmed/22859988
- McDowell, A. et al. The opportunistic pathogen Propionibacterium acnes: Insights into typing, human disease, clonal diversification and CAMP factor evolution. PLoS ONE 8, e70897 (2013). https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0070897
- Fitz-Gibbon, S. et al. Propionibacterium acnes strain populations in the human skin microbiome associated with acne. J Investig Dermatol 133, 2152 – 2160 (2013). https://www.ncbi.nlm.nih.gov/pubmed/23337890
- Valanne, S. et al. CAMP factor homologues in Propionibacterium acnes: a new protein family differentially expressed by types I and II. Microbiology 151, 1369 – 1379 (2005). https://www.ncbi.nlm.nih.gov/pubmed/15870447
- Holland, C. et al. Proteomic identification of secreted proteins of Propionibacterium acnes. BMC Microbiol 10, 230 (2010). https://www.ncbi.nlm.nih.gov/pubmed/20799957
- Jasson, F. et al. Different strains of Propionibacterium acnes modulate differently the cutaneous innate immunity. Exp Dermatol 22, 587 – 592 (2013). https://www.ncbi.nlm.nih.gov/pubmed/23947673
- Kuehnast, T., Cakar, F., Weinhäupl, T., Pilz, A., Selak, S., Schmidt, M. A., Rüter, C. & Schild, S. Comparative analyses of biofilm formation among different Cutibacterium acnes isolates. Int J Med Microbiol 308, 1027-1035 (2018). https://www.ncbi.nlm.nih.gov/pubmed/30268774
- Hazarika, N. Acne vulgaris: new evidence in pathogenesis and future modalities of treatment. J Dermatolog Treat 32, 277-285 (2021). https://pubmed.ncbi.nlm.nih.gov/31393195/
- Dréno, B., Pécastaings, S., Corvec, S., Veraldi, S., Khammari, A. & Roques, C. Cutibacterium acnes (Propionibacterium acnes) and acne vulgaris: a brief look at the latest updates. J Eur Acad Dermatol Venereol 32 Suppl 2, 5-14 (2018). https://www.ncbi.nlm.nih.gov/pubmed/29894579
- Dagnelie, M. A., Montassier, E., Khammari, A., Mounier, C., Corvec, S. & Dréno, B. Inflammatory skin is associated with changes in the skin microbiota composition on the back of severe acne patients. Exp Dermatol 28, 961-967 (2019). https://pubmed.ncbi.nlm.nih.gov/31173650/
- Huang, T. Y., Jiang, Y. E. & Scott, D. A. Culturable bacteria in the entire acne lesion and short-chain fatty acid metabolites of Cutibacterium acnes and Staphylococcus epidermidis isolates. Biochem Biophys Res Commun 622, 45-49 (2022). https://pubmed.ncbi.nlm.nih.gov/35843093/
 Acne.org Products
Acne.org Products